Approximately 1 percent of human DNA is made of protein-coding genes, while the remaining 99 percent comprises non-coding genes. Non-coding DNA does not provide instructions for making proteins and was previously thought to be “junk” by scientists.
Long non-coding RNAs (lncRNAs) are a type of RNA with lengths exceeding 200 nucleotides that are not translated into protein. lncRNAs can be significant components in epigenetic processes that function to regulate gene expression at the transcriptional and post-transcriptional levels. The role of lncRNAs in human Mendelian disease on a large scale remains unknown.
For the past 20 years, geneticists haven’t agreed on the number of protein-coding genes. The latest estimates put the figure around 20,000, including the almost 5,000 genes that were not uncovered before.
Read the original publication of this study here:[ Non-coding deletions identify Maenli lncRNA as a limb-specific En1 regulator ]
This study aimed to show that genetic ablation of a lncRNA locus on human chromosome 2 causes a severe congenital limb malformation.
Non-coding deletions identify Maenli lncRNA as a limb-specific En1 regulator
An international team of researchers uncovered the mechanism behind a rare genetic disease that involves non-coding DNA sequences.
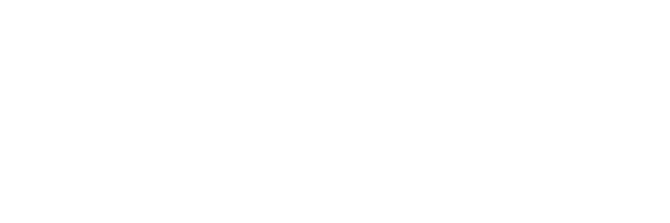
In this study, researchers identified homozygous 27–63-kilobase deletions located 300 kilobases upstream of the engrailed-1 gene (EN1) in patients with complex limb malformation resulting in disproportionately short middle parts of the limbs (mesomelia), two or more digits are fused (syndactyly), and ventral nails.
The En1 gene has been known to regulate multicellular organisms’ development, specifically in the dorsal midbrain and anterior hindbrain. Re-engineering these human deletions through deactivation of its transcription in mice resulted in a complete loss of En1 expression in the limb and a double dorsal-limb phenotype that resembles the similar piece of non-coding DNA missing in the patients.
Further genome-wide analysis in the developing mouse limb showed a four-exon-long non-coding transcript within the deleted region, which was previously unknown. This RNA molecule is named Maenli, which stands for Master Activator of Engrailed-1 in the Limb.
Functional dissection of the Maenli locus showed that its transcriptional activity is required for limb-specific En1 cis activation, thereby fine-tuning the gene-regulatory networks controlling dorsoventral polarity in the developing limb bud.
The research findings demonstrate that mutations involving lncRNA loci can result in human Mendelian disease.
Takeaways:
- Researchers uncovered a previously unknown genome sequence by reengineering the human deletions of homozygous 27–63-kilobase located 300 kilobases upstream of EN1 in mice.
- Genetic ablation of a lncRNA locus on human chromosome 2 causes a severe congenital limb malformation
- Mutations involving long non-coding RNAs (lncRNA) loci possibly result in Mendelian disease.
You can read the original publication of this study here: [Non-coding deletions identify Maenli lncRNA as a limb-specific En1 regulator ]