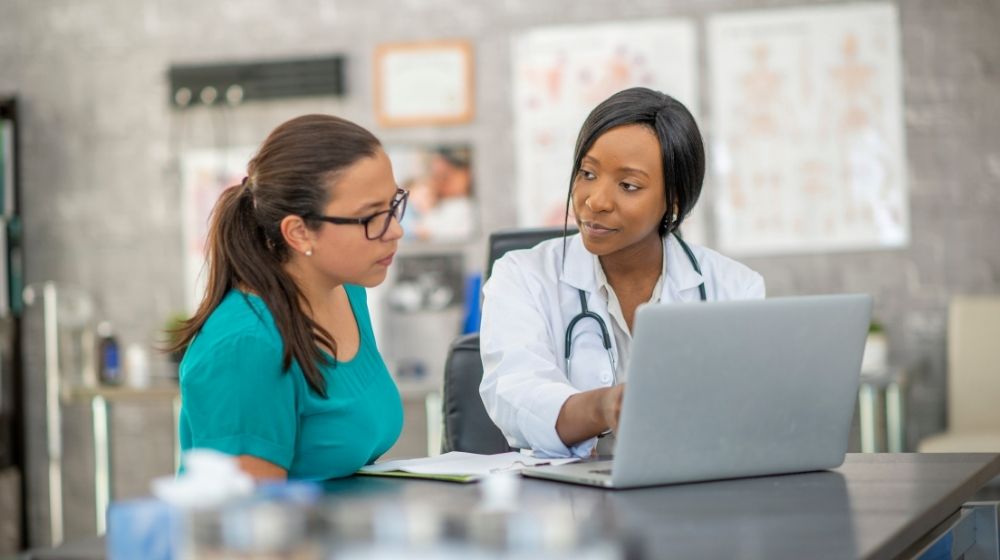
Using a new algorithm, researchers reveal a previously undetected collection of cancer driver genes. As a result, our knowledge of epigenetic mechanisms in tumorigenesis and cancer prediction is improving.
RELATED: Machine Learning and Clinical Epigenetics
In this article:
Cancer Driver Genes
Cancers stem from a build-up of genetic changes in the DNA of normal cells. These changes upset the balance between cell division and apoptosis (programmed cell death) and result in a normal cell becoming a cancer cell. But, not all abnormalities lead to cancer. Some may not contribute at all. The terms driver and passenger mutation embody this concept.
There’s a causal connection between a driver mutation and cells becoming cancerous. Identifying driver mutations and the cancer genes they change is a central aim of cancer research.
Genes with driver mutations give a growth advantage to the cancer cell. Their cancer progression functions classify them as oncogenes (OGs) and tumor suppressor genes (TSGs).
However, a passenger mutation doesn’t give a clonal growth advantage. Therefore they have not contributed to cancer development.
These harmless passenger mutations often occur during cell division. Thus, cells with driver mutations may already have biologically inert mutations within their genome, as they were previously a carrier of passenger mutations before the final driver mutation tipped them over to become cancerous.
As a result, passenger mutations are often also present in cancer cells.
Cancer driver genes that don’t mutate frequently are often identical to passenger genes with chance mutations in genome sequencing data. So, to discern driver mutations from passenger mutations is one of the top challenges in genome analysis.
Previous studies relied on various non-mutational datasets to identify driver gene mutations. This restricts identification to the cancer genome elements, a small number of cancer samples, or a subset of the available mutational classes.
Furthermore, detection methods based on the mutational background model or functional impact model can hardly spot infrequently mutated cancer driver genes. However, joining epigenetic, genomic, and phenotypic data may pinpoint these genes.
DORGE Algorithm
DORGE (Discovery of Oncogenes and tumor suppressor genes using Genetic and Epigenetic features) is the latest prediction algorithm from the UCI School of Medicine.
The algorithm combines public data on epigenetic and genetic variations to refine the prediction of cancer driver genes. It includes two prediction algorithms: DORGE-TSG and DORGE-OG; both algorithms use a type of machine learning trained on neutral genes (NGs) and Cancer Gene Census (CGC).
During a recent study, DORGE discovered histone changes as powerful predictors for TSGs, and it found missense mutations, methylation differences, and super-enhancers as strong predictors for OGs. Researchers also found dual-functional genes are enriched at hubs in drug/compound-gene networks and protein-protein interaction (PPI), which DORGE predicted as both TSGs and OGs.
Additionally, DORGE predictions were widely verified using objective functional genomics data. DORGE performed the best in terms of sensitivity, precision, and overall accuracy compared to existing predictive analytics algorithms for cancer driver gene prediction.
RELATED: We Can Now Turn Back the Biological Clock: 5 Advantages of Epigenetics
How Cancer Prediction Is Improving
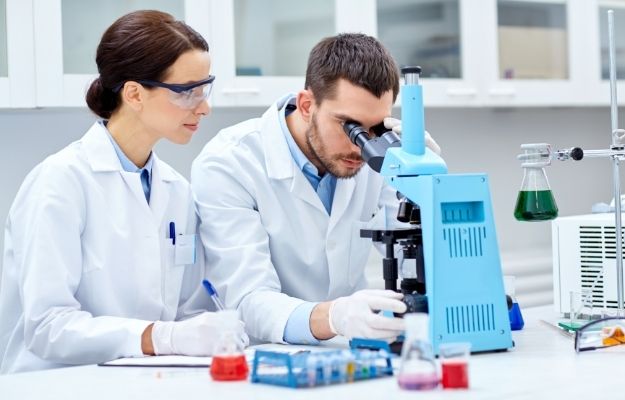

Data-driven discovery of cancer driver genes, including oncogenes (OGs) and tumor suppressor genes (TSGs), is crucial for cancer prevention, diagnosis, and treatment.
According to Wei Li, Prof. of bioinformatics in the Department of Biological Chemistry at the UCI School of Medicine, existing bioinformatics algorithms don’t use epigenetic features to sufficiently predict cancer driver genes. Instead, their algorithm uses public data to discover the epigenetic changes that play significant roles in cancer driver gene disruption.
Researchers were able to identify genes with rare mutations because DORGE classifies TSGs and OGs by combining extensive genetic and epigenetic data. The ability to predict TSGs and OGs separately lets DORGE identify novel dual-functional cancer driver genes.
When using epigenetics, DORGE is unbiased in gene selection to predict cancer driver genes, one of the shortcomings in existing studies.
Above all, cancer prediction is improving because the cancer driver genes, predicted by DORGE, included both known cancer driver genes and driver genes not listed in current research.
To conclude, integrating epigenetic data results in more complete cancer driver gene predictions. Moreover, DORGE will be vital for cancer biology, especially for creating targeted therapies and personalized cancer treatment.
If you’re interested in learning more about epigenetics and research developments, visit the TruDiagnostic website today.
Do you think cancer prediction is improving? What other developments would you like to see? Share your thoughts in the comments section!
Source:
Up Next: